postProcess/VideoAxi.py
#!/usr/bin/env python3
# -*- coding: utf-8 -*-Basilisk C Simulation Data Post-Processor and Visualizer
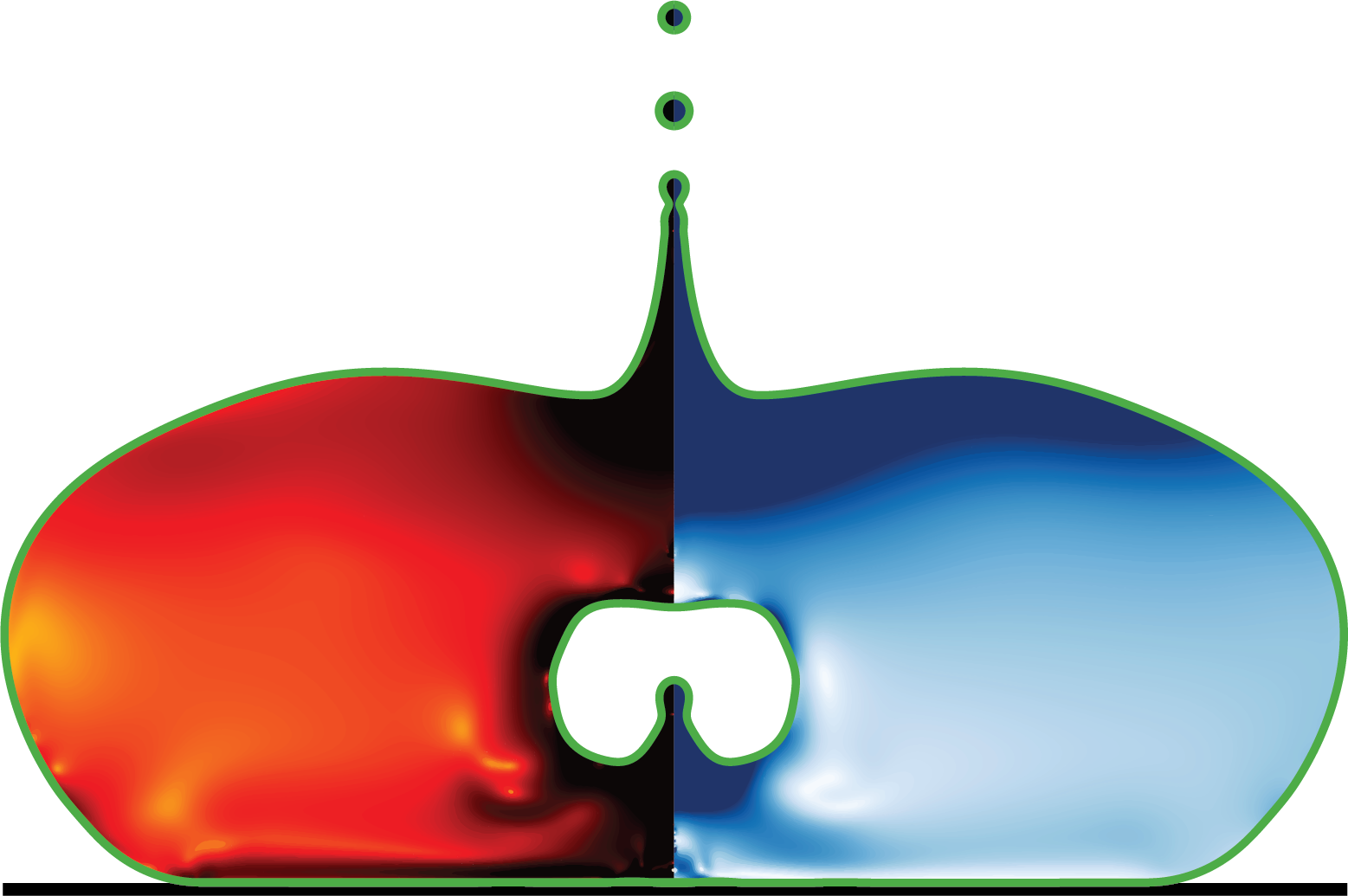
This script processes time-series data from Basilisk C multiphase flow simulations, generating high-quality visualizations of interfacial dynamics and flow fields. It creates contour plots showing strain rate tensor magnitude (D2), velocity fields, and viscoelastic stress traces for bubble/drop dynamics studies.
The script supports parallel processing for efficient handling of large time-series datasets and produces publication-quality figures with proper LaTeX formatting.
Usage: python viz_simulation.py [–CPUs 8] [–nGFS 550] [–ZMAX 4.0] [–RMAX 2.0] [–ZMIN -4.0]
Dependencies: - numpy: Numerical array operations - matplotlib: Plotting and visualization - subprocess: External executable communication - multiprocessing: Parallel processing support - argparse: Command-line argument parsing
External Dependencies: - ./getFacet2D: Basilisk executable for interface extraction - ./getData-elastic-scalar2D: Basilisk executable for field data extraction
Author: Vatsal Sanjay Contact: [email protected] Affiliation: Physics of Fluids Group Last updated: Jul 24, 2024
import numpy as np
import os
import subprocess as sp
import matplotlib
import matplotlib.pyplot as plt
from matplotlib.collections import LineCollection
from matplotlib.ticker import StrMethodFormatter
import multiprocessing as mp
from functools import partial
import argparse
import matplotlib.colors as mcolors
# ===============================
# Configuration and Settings
# ===============================
# Custom colormap for viscoelastic stress visualization
custom_colors = ["white", "#DA8A67", "#A0522D", "#400000"]
CUSTOM_CMAP = mcolors.LinearSegmentedColormap.from_list("custom_hot", custom_colors)
# Global matplotlib settings for publication-quality output
matplotlib.rcParams['font.family'] = 'serif'
matplotlib.rcParams['text.usetex'] = True
matplotlib.rcParams['text.latex.preamble'] = r'\usepackage{amsmath}'
# Default visualization parameters
DEFAULT_CONFIG = {
'grids_per_r': 128, # Grid resolution factor
'line_width': 2, # Interface line width
'axes_label_size': 50, # Font size for axis labels
'tick_label_size': 20, # Font size for tick labels
'interface_line_width': 4, # Line width for interface boundaries
'strain_vmax': 2.0, # Maximum strain rate for colorbar
'strain_vmin': -3.0, # Minimum strain rate for colorbar
'stress_vmax': 2.0, # Maximum stress trace for colorbar
'stress_vmin': -3.0, # Minimum stress trace for colorbar
}
# ===============================
# Interface Extraction Functions
# ===============================
def gettingFacets(filename, includeCoat='true'):Extract interface facets from Basilisk simulation data.
This function communicates with the Basilisk getFacet2D executable to extract interface polygons from simulation snapshots. It handles both axisymmetric coordinates (r,z) and creates symmetric segments for visualization.
Args: filename (str): Path to the Basilisk snapshot file includeCoat (str): Whether to include coating layer in extraction. Accepts ‘true’ or ‘false’. Defaults to ‘true’.
Returns: list: List of line segments [(start_point, end_point)] for visualization. Each segment is a tuple ((r1, z1), (r2, z2)) representing interface boundaries.
Raises: subprocess.CalledProcessError: If the getFacet2D executable fails FileNotFoundError: If the specified snapshot file doesn’t exist
Note: The function assumes axisymmetric geometry and creates symmetric segments by mirroring across r=0. This is typical for bubble/drop simulations.
exe = ["./getFacet2D", filename, includeCoat]
try:
p = sp.Popen(exe, stdout=sp.PIPE, stderr=sp.PIPE)
stdout, stderr = p.communicate()
except FileNotFoundError:
raise FileNotFoundError(f"getFacet2D executable not found. Ensure it's compiled and in the current directory.")
temp1 = stderr.decode("utf-8")
temp2 = temp1.split("\n")
segs = []
skip = False
# Process interface data - skip empty lines and extract coordinate pairs
if len(temp2) > 1e2: # Ensure we have sufficient data points
for n1 in range(len(temp2)):
temp3 = temp2[n1].split(" ")
if temp3 == ['']:
skip = False
pass
else:
if not skip:
# Extract paired coordinate points
temp4 = temp2[n1+1].split(" ")
r1, z1 = np.array([float(temp3[1]), float(temp3[0])])
r2, z2 = np.array([float(temp4[1]), float(temp4[0])])
# Add symmetric segments for axisymmetric visualization
segs.append(((r1, z1), (r2, z2)))
segs.append(((-r1, z1), (-r2, z2)))
skip = True
return segs
def gettingfield(filename, zmin, zmax, rmax, nr):Extract field data from Basilisk simulation snapshots.
Communicates with the getData-elastic-scalar2D executable to extract flow field variables including strain rate, velocity, and stress tensors on a uniform grid for visualization.
Args: filename (str): Path to the Basilisk snapshot file zmin (float): Minimum z-coordinate for data extraction zmax (float): Maximum z-coordinate for data extraction rmax (float): Maximum r-coordinate for data extraction nr (int): Number of grid points in radial direction
Returns: tuple: (R, Z, D2, vel, taup, nz) where: - R (numpy.ndarray): Radial coordinate mesh (nz x nr) - Z (numpy.ndarray): Axial coordinate mesh (nz x nr) - D2 (numpy.ndarray): Strain rate tensor magnitude (nz x nr) - vel (numpy.ndarray): Velocity magnitude (nz x nr) - taup (numpy.ndarray): Stress tensor trace (nz x nr) - nz (int): Number of grid points in axial direction
Raises: subprocess.CalledProcessError: If the getData-elastic-scalar2D executable fails ValueError: If the extracted data dimensions are inconsistent
Note: The function automatically determines nz based on the total data points and the specified nr. All returned arrays are reshaped to 2D meshgrids.
exe = ["./getData-elastic-scalar2D", filename, str(zmin), str(0), str(zmax), str(rmax), str(nr)]
try:
p = sp.Popen(exe, stdout=sp.PIPE, stderr=sp.PIPE)
stdout, stderr = p.communicate()
except FileNotFoundError:
raise FileNotFoundError(f"getData-elastic-scalar2D executable not found. Ensure it's compiled and in the current directory.")
temp1 = stderr.decode("utf-8")
temp2 = temp1.split("\n")
# Initialize temporary lists for field data
Rtemp, Ztemp, D2temp, veltemp, taupTemp = [], [], [], [], []
# Parse field data from executable output
for n1 in range(len(temp2)):
temp3 = temp2[n1].split(" ")
if temp3 == ['']:
pass
else:
Ztemp.append(float(temp3[0]))
Rtemp.append(float(temp3[1]))
D2temp.append(float(temp3[2]))
veltemp.append(float(temp3[3]))
taupTemp.append(float(temp3[4]))
# Convert to numpy arrays and reshape to 2D mesh
R = np.asarray(Rtemp)
Z = np.asarray(Ztemp)
D2 = np.asarray(D2temp)
vel = np.asarray(veltemp)
taup = np.asarray(taupTemp)
# Calculate number of axial grid points
nz = int(len(Z)/nr)
print(f"Grid dimensions: nr={nr}, nz={nz}")
# Reshape arrays to 2D mesh format
R.resize((nz, nr))
Z.resize((nz, nr))
D2.resize((nz, nr))
vel.resize((nz, nr))
taup.resize((nz, nr))
return R, Z, D2, vel, taup, nz
# ===============================
# Visualization Functions
# ===============================
def process_timestep(ti, folder, nGFS, GridsPerR, rmin, rmax, zmin, zmax, lw):Process and visualize a single simulation timestep.
This function handles the complete visualization pipeline for one timestep: extracting interface data, field data, creating dual-sided contour plots with colorbars, and saving the result as a high-resolution image.
Args: ti (int): Timestep index (will be converted to simulation time) folder (str): Output directory for saved images nGFS (int): Total number of simulation files (used for validation) GridsPerR (int): Grid resolution parameter (grids per unit radius) rmin (float): Minimum radial coordinate for plotting rmax (float): Maximum radial coordinate for plotting zmin (float): Minimum axial coordinate for plotting zmax (float): Maximum axial coordinate for plotting lw (float): Line width for boundary boxes
Returns: None: Function saves visualization directly to file
Note: - Creates symmetric visualization about r=0 axis - Uses logarithmic scaling for strain rate and stress data - Implements custom colormap for viscoelastic stress fields - Handles missing files gracefully with informative error messages
# Convert timestep index to simulation time
t = 0.01 * ti
place = f"intermediate/snapshot-{t:.4f}"
name = f"{folder}/{int(t*1000):08d}.png"
# Check if input file exists
if not os.path.exists(place):
print(f"{place} File not found!")
return
# Skip if output already exists
if os.path.exists(name):
print(f"{name} Image present!")
return
# Extract interface data with and without coating
segs1 = gettingFacets(place) # With coating
segs2 = gettingFacets(place, 'false') # Without coating
# Validate interface data
if not segs1 and not segs2:
print(f"Problem in the available file {place}")
return
# Extract field data on uniform grid
nr = int(GridsPerR * rmax)
R, Z, taus, vel, taup, nz = gettingfield(place, zmin, zmax, rmax, nr)
zminp, zmaxp, rminp, rmaxp = Z.min(), Z.max(), R.min(), R.max()
# ===============================
# Create Visualization
# ===============================
AxesLabel, TickLabel = DEFAULT_CONFIG['axes_label_size'], DEFAULT_CONFIG['tick_label_size']
fig, ax = plt.subplots()
fig.set_size_inches(19.20, 10.80) # High-resolution output
# Draw boundary box and symmetry axis
ax.plot([0, 0], [zmin, zmax], '-.', color='grey', linewidth=lw) # Symmetry axis
ax.plot([rmin, rmin], [zmin, zmax], '-', color='black', linewidth=lw)
ax.plot([rmin, rmax], [zmin, zmin], '-', color='black', linewidth=lw)
ax.plot([rmin, rmax], [zmax, zmax], '-', color='black', linewidth=lw)
ax.plot([rmax, rmax], [zmin, zmax], '-', color='black', linewidth=lw)
# Add interface lines
line_segments = LineCollection(segs2, linewidths=DEFAULT_CONFIG['interface_line_width'],
colors='green', linestyle='solid')
ax.add_collection(line_segments)
line_segments = LineCollection(segs1, linewidths=DEFAULT_CONFIG['interface_line_width'],
colors='blue', linestyle='solid')
ax.add_collection(line_segments)
# Create strain rate contour plot (left side, mirrored)
cntrl1 = ax.imshow(taus, cmap="hot_r", interpolation='Bilinear', origin='lower',
extent=[-rminp, -rmaxp, zminp, zmaxp],
vmax=DEFAULT_CONFIG['strain_vmax'], vmin=DEFAULT_CONFIG['strain_vmin'])
# Create stress trace contour plot (right side)
cntrl2 = ax.imshow(taup, interpolation='Bilinear', cmap=CUSTOM_CMAP, origin='lower',
extent=[rminp, rmaxp, zminp, zmaxp],
vmax=DEFAULT_CONFIG['stress_vmax'], vmin=DEFAULT_CONFIG['stress_vmin'])
# Set plot properties
ax.set_aspect('equal')
ax.set_xlim(rmin, rmax)
ax.set_ylim(zmin, zmax)
ax.set_title(f'$t/\\tau_\\gamma$ = {t:4.3f}', fontsize=TickLabel)
# Add colorbars
l, b, w, h = ax.get_position().bounds
# Left colorbar for strain rate
cb1 = fig.add_axes([l-0.04, b, 0.03, h])
c1 = plt.colorbar(cntrl1, cax=cb1, orientation='vertical')
c1.set_label(r'$\log_{10}\left(\|\mathcal{D}\|\right)$', fontsize=TickLabel, labelpad=5)
c1.ax.tick_params(labelsize=TickLabel)
c1.ax.yaxis.set_ticks_position('left')
c1.ax.yaxis.set_label_position('left')
c1.ax.yaxis.set_major_formatter(StrMethodFormatter('{x:,.1f}'))
# Right colorbar for stress trace
cb2 = fig.add_axes([l+w+0.01, b, 0.03, h])
c2 = plt.colorbar(cntrl2, cax=cb2, orientation='vertical')
c2.ax.tick_params(labelsize=TickLabel)
c2.set_label(r'$\log_{10}\left(\text{tr}\left(\mathcal{A}\right)-1\right)$', fontsize=TickLabel)
c2.ax.yaxis.set_major_formatter(StrMethodFormatter('{x:,.2f}'))
ax.axis('off') # Remove axis ticks and labels for cleaner look
# Save high-quality output
plt.savefig(name, bbox_inches="tight", dpi=300)
plt.close() # Free memory
# ===============================
# Main Execution Function
# ===============================
def main():Main function that orchestrates the parallel processing of simulation data.
Sets up command-line argument parsing, configures multiprocessing pool, and coordinates the visualization of multiple timesteps in parallel. Uses all available CPU cores by default for optimal performance.
Command-line Arguments: –CPUs (int): Number of CPU cores to use (default: all available) –nGFS (int): Number of simulation timesteps to process (default: 550) –ZMAX (float): Maximum axial coordinate (default: 4.0) –RMAX (float): Maximum radial coordinate (default: 2.0) –ZMIN (float): Minimum axial coordinate (default: -4.0)
Returns: None: Creates output directory and processes all timesteps
Example: # Process 1000 timesteps using 16 CPU cores python viz_simulation.py –CPUs 16 –nGFS 1000 –ZMAX 5.0
Note: The output directory ‘Video’ is created automatically if it doesn’t exist. Each timestep is processed independently, allowing for efficient parallel execution.
# Set up command-line argument parsing
parser = argparse.ArgumentParser(description="Process Basilisk simulation data for visualization")
parser.add_argument('--CPUs', type=int, default=mp.cpu_count(),
help='Number of CPUs to use (default: all available)')
parser.add_argument('--nGFS', type=int, default=550,
help='Number of restart files to process (default: 550)')
parser.add_argument('--ZMAX', type=float, default=4.0,
help='Maximum Z value (default: 4.0)')
parser.add_argument('--RMAX', type=float, default=2.0,
help='Maximum R value (default: 2.0)')
parser.add_argument('--ZMIN', type=float, default=-4.0,
help='Minimum Z value (default: -4.0)')
args = parser.parse_args()
# Extract parameters
CPUStoUse = args.CPUs
nGFS = args.nGFS
ZMAX = args.ZMAX
RMAX = args.RMAX
ZMIN = args.ZMIN
# Set processing parameters
num_processes = CPUStoUse
rmin, rmax, zmin, zmax = [-RMAX, RMAX, ZMIN, ZMAX]
GridsPerR = DEFAULT_CONFIG['grids_per_r']
lw = DEFAULT_CONFIG['line_width']
folder = 'Video'
# Create output directory
if not os.path.isdir(folder):
os.makedirs(folder)
print(f"Created output directory: {folder}")
print(f"Starting visualization process with {num_processes} CPUs...")
print(f"Processing {nGFS} timesteps from {ZMIN} to {ZMAX} in Z and {-RMAX} to {RMAX} in R")
# Create multiprocessing pool and process all timesteps
with mp.Pool(processes=num_processes) as pool:
# Create partial function with fixed arguments
process_func = partial(process_timestep,
folder=folder, nGFS=nGFS,
GridsPerR=GridsPerR, rmin=rmin, rmax=rmax,
zmin=zmin, zmax=zmax, lw=lw)
# Map the processing function to all timesteps
timesteps = list(range(nGFS))
pool.map(process_func, timesteps)
print(f"Visualization complete! Images saved in {folder}/")
if __name__ == "__main__":
main()